 username@email.com
username@email.com
In this lesson, we will review how eukaryotic genes are regulated, which enzymes are expressed by genes, and how this allows cells to function across many conditions and environments.
Eukaryotic cells face the same problems as prokaryotic cells. However, eukaryotic cells (except for the protists) are highly differentiated and specialized within the organism. In order to achieve this high level of specialization, different genes must be turned on or off in different cells. A highly specialized cell, such as a neuron, might express only 3 to 5 percent of the genes in its nucleus. A different type of cell will express a different 3 to 5 percent. All cells need to perform certain functions, so there are genes that are always “turned on.” However, as cells become more specialized during human development, the number of active genes in any given cell decreases.

Eukaryotes need to regulate their genes for different reasons than prokaryotes. In prokaryotes, gene regulation allowed them to respond to their environment efficiently and economically. While eukaryotes can respond to their environment, the environment of most eukaryotic cells in complex organisms is relatively stable. There are many regulatory systems, for example, in the human body, where the function is to maintain a stable environment for the cells. The main reason eukaryotes need to regulate their genes is cell specialization. Prokaryotes are relatively simple unicellular organisms, while eukaryotes may have hundreds of different cell types, each differentiated to serve a specialized function. Each cell type differentiates by activating a different subset of genes. Because of the multitude of cell types, the regulation of gene expression required to bring about such differentiation is necessarily complex. One way this complexity is demonstrated is in multiple levels of gene expression regulation. The latest estimates are that a human cell, which is a eukaryotic cell, contains 20,000 to 25,000 active genes — at least 10 times that of a prokaryotic cell.
Some genes are expressed in all eukaryote cells at all times. These genes are responsible for the routine metabolic functions (such as respiration) common to all cells. Some genes are expressed as a cell enters a particular pathway of differentiation. Some genes are expressed all the time in only those cells that have differentiated in a particular way. For example, a plasma cell expresses continuously the genes for the antibody it synthesizes. Some genes are expressed only as conditions change in and around the cell. For example, the presence or absence of a hormone may turn on, or off, certain genes in that cell.
How many genes are estimated to be in each human cell?
C is the correct answer. The latest research shows that human cells have about 25,000 active genes.
There is about 1,000 times as much DNA in a human cell as in an E. coli cell, but only about 10 — 20 times as many genes. How can this be?
A is the correct answer. Most of the human DNA strand consists of non-coding introns. Choice B is incorrect because DNA packaging is simpler in a prokaryotic cell. Choice C is incorrect; although the genes in specialized cells may be inactivated, they are still present. Choice D is incorrect because E. coli is able to respond readily to its environment.
There are several methods used by eukaryotes to regulate genes. The primary method is by altering the rate of transcription of the gene. Secondary methods include:

Where in the cell does gene transcription take place?
The correct answer is D, because transcription is the process of writing the code to an mRNA molecule. Choice A is incorrect; although mitochondria contain genetic information, they are separate from nuclear genetic information. Choice B is not correct because the cytoplasm only serves to transport mRNA to the ribosome. Choice C is not correct because the ribosome is the location of translation, not transcription.
Regulation of gene transcription is the most important means by which gene expression is regulated in eukaryotes, as it was in prokaryotes. However, there are important differences between transcriptional regulation in prokaryotes and eukaryotes. Because of the complexity of eukaryotic patterns of gene expression, each eukaryotic gene is controlled by a separate promoter. Unlike prokaryotes, eukaryotic genes are not organized into operons. Prokaryotic genes are regulated primarily by repressors while eukaryotic genes are primarily regulated by transcriptional activators. These activators are called transcription factors.

Which of the following is the primary method of regulating gene expression in eukaryotes?
A is the correct answer. Regulation of gene expression is accomplished through changing the rate of transcription. Choices B, C, and D are methods of lesser importance.
After transcription, the RNA must be processed before it can be translated. RNA processing involves the addition of a 5′ cap, addition of a 3′ poly (A) tail, and removal of introns. Regulation of gene expression can also occur during RNA processing in two different ways: whether an RNA gets processed at all, and which exons are retained in the mRNA (or which introns are excised).
The first type of regulation can determine whether or not an mRNA gets translated. If RNA is not processed, it will not be transported out of the nucleus, and will not be translated.
The second type of regulation can affect the function of the protein produced. Some genes have exons that can be arranged in several different orders, a process called exon shuffling. When the exons are rearranged, the polypeptide produced could have a different function. For example, in mammals the calcitonin gene produces a hormone in one cell type and a neurotransmitter in another cell type, because of exon shuffling.

Molecules of mRNA can usually produce many copies of a polypeptide before becoming degraded. If a molecule lasts for only a short time, it will produce only a few copies of the polypeptide. A different mRNA molecule might last an hour or more; in the process producing thousands of its polypeptide molecule.
Prokaryote mRNA typically only lasts for a few minutes, which is one reason bacteria and other prokaryotes can respond so rapidly to environmental changes by varying their protein synthesis. Eukaryote mRNA typically lasts much longer; hours, days, or even weeks.
The mRNAs for the hemoglobin polypeptides (alpha-globin and beta-globin) found in developing human red blood cells are unusually stable. This is the principle behind the regulation of mRNA longevity. mRNAs from different genes have their approximate lifespan encoded in them. This serves to help regulate how much of each polypeptide is produced.
The information for lifespan is found in the untranslated trailer region (UTR) at the 3′ end. The sequence AUUUA, when found in the 3′ UTR, is a signal for early degradation (and therefore a short lifespan). The more times the sequence is present, the shorter the lifespan of the mRNA. Because the approximate lifespan of a strand of mRNA is encoded in the nucleotide sequence, this is a set property of each different mRNA; the longevity of an mRNA cannot be varied.
Translation of the mRNA molecule can also be regulated. Translation control mechanisms typically block the initiation stage of polypeptide synthesis. Regulatory proteins can bind to specific sequences within the leader region and the 5′ end of the mRNA strand, effectively preventing ribosomes from attaching. Global control of all or most of the translation of mRNA can be initiated by protein factors. For example, if hemes are in short supply in a developing red blood cell for some reason, a regulatory protein can inactivate a translation initiation factor by phosphorylating it. This halts all translation, but mainly affects the translation of hemoglobin mRNA since the developing red blood cell is primarily engaged in hemoglobin production.
Global control of translation is important during development of embryos. For example, many animals sequester large amounts of mRNA in their eggs, and those mRNA molecules are not translated unless the egg is fertilized. At that point, translation is triggered by the sudden activation of translation initiation factors, resulting in a burst of protein synthesis.
After translation, there is one more opportunity for a cell to regulate gene expression — through protein degradation. Polypeptides must be further processed to become functional proteins. For example, the protein that will become insulin must be cleaved before it is functional. Other proteins acquire sugars, and phosphate groups activate or deactivate regulatory proteins. Polypeptides frequently must be transported to other parts of the cell. All of these processes could serve as a mechanism for regulation.
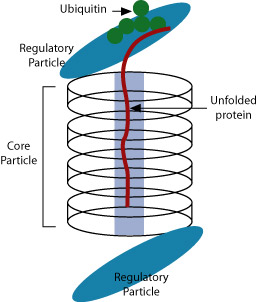
Proteins marked for degradation are tagged with a small bit of protein called ubiquitin. It is not clear how proteins are chosen for destruction. Once marked, the tagged proteins are recognized by a large structure called a proteasome. The proteasome unfolds the tagged protein, sequesters it in its interior, and chops it into small peptide bits that are recycled. The ubiquitin and proteasome are also recycled.
In prokaryotes, genes that work together to produce a set of related enzymes are grouped together into an operon. Eukaryotic cells do not have operons, so they require some other mechanism for controlling several genes at the same time. This coordinated gene expression probably depends on a specific control element or set of elements that are associated with every gene in a dispersed group. A single key sets all of the genes to turn off or on at the same time.
In prokaryotes, an example of this sort of coordinated gene expression is the activation of a variety of genes by a steroid hormone. These steroid hormones (sex hormones are one example) have a variety of effects on the body. A steroid hormone acts like a chemical signal, entering each cell and binding to a specific receptor protein in the cell. When turned on, these receptors act as transcription activators. Every gene that will be turned on by that hormone has a control element that recognizes the transcription activator.
Genes may be turned on in one type of cell and not in another, depending on whether the transcription factor for the gene’s enhancer is active or not. For example, consider a cell in muscle tissue and a cell in skin tissue from the same person. Both have the exact same set of genes. In the muscle tissue cell, a particular set of transcription factors has become active causing the set of genes needed by muscle cells to be expressed. In the epidermal tissue cell, a different set of transcription factors is active, causing a different set of genes to be expressed.
How does the body know which transcription factors to send to which cell types and how do they get there? This is a difficult question with a very complicated answer. Many of the details have not yet been worked out by biologists. However, there are a number of mechanisms by which transcription factors can be regulated.
Steroid hormones, such as testosterone and progesterone, are derived from lipids (specifically cholesterol). Because they are derived from lipids, they can easily pass through the cytoplasmic membrane into a cell, where they bind to their specific receptor. Steroid receptors are transcription factors that become activated when they bind to their ligand. Once activated, they initiate transcription of a specific set of genes.

Since peptide hormones cannot easily cross the cytoplasmic membrane, their receptors are found on the surface of the cell. When bound to its ligand, these receptors initiate a complex series of biochemical reactions inside the cell, with the ultimate result being the activation of a transcription factor (often by phosphorylation, which initiates transcription of a specific set of genes.

Why are human muscle cells and bone cells different?
C is the correct answer. Only 2 to 5 percent of the genes in a specialized cell is activated. Choice A is incorrect because all of the gene sets are the same. Choice B is incorrect because eukaryotic cells are not known to contain operons. Choice D is incorrect because each specific gene produces a specific protein unless the mRNA is damaged or reshuffled.